First time using Scellnetor or need help??
Check out our documentation here.
Or check out our short screencast →→→Scellnetor
- Upload your data or try a test set
- Select the template-plot you want to use
- Draw trajectories or pick clusters
- Choose parameters for the constrained hierarchical clustering algorithm
- Inspect and download results
You can upload your own AnnData object as a H5AD file.
You can upload your old data here as a ZIP file.
Tutorials
Alternatively, you can try some of the premade examples to give scellnetor a spin! The test dataset are designed for easy and quick overview of the features of Scellnetor!

Try single cells from a study on peripheral blood mononuclear cells
The data is based on: Clustering 3K PBMCs
Try single cells from another hematopoiesis study
The data is based on: DPT for hematopoiesis in mouse (Moignard et al., 2015)
Try single cells from a study on CD8+ T-cells in lung cancer
Test data is based on: Guo X et al. (2018)
hej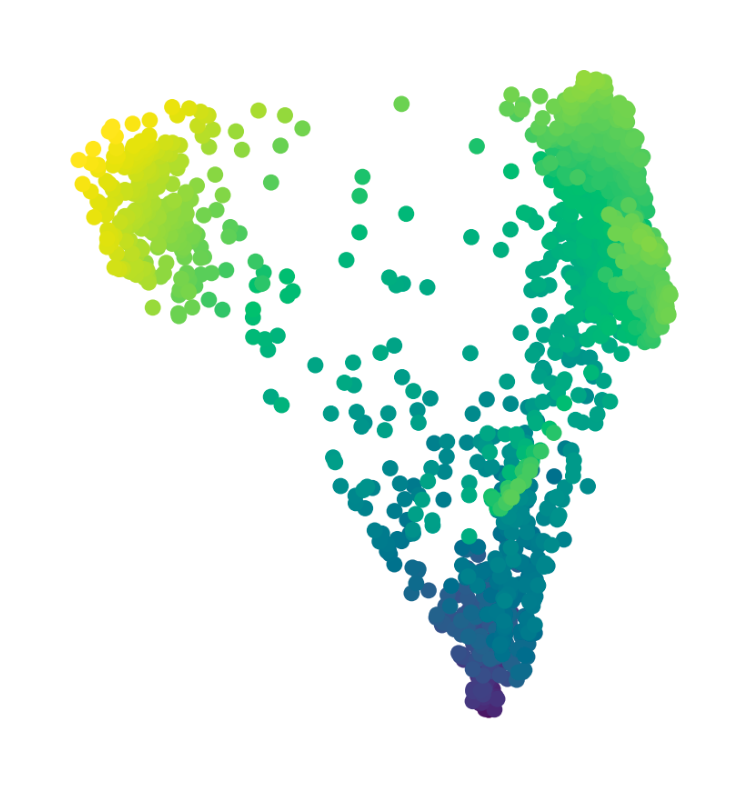
Try single cells from a study on dysfunctional T-cells in chronic infections
The data is based on: Kanev et al. (2019)